-Search query
-Search result
Showing 1 - 50 of 106 items for (author: ahn & hm)
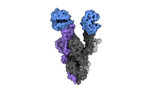
EMDB-40976: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to two mink ACE2 receptors
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B
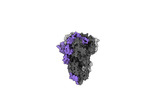
EMDB-40977: 
Cryo-EM structure of mink variant Y453F trimeric spike protein
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

EMDB-40978: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to one mink ACE2 receptors at downRBD conformation
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

EMDB-40979: 
Cryo-EM structure of the RBD-ACE2 interface of the SARS-CoV-2 trimeric spike protein bound to ACE2 receptor after local refinement at upRBD conformation
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

EMDB-40980: 
Cryo-EM structure of the RBD-ACE2 interface of the SARS-CoV-2 trimeric spike protein bound to ACE2 receptor after local refinement at downRBD conformation.
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B
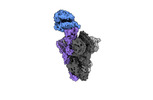
EMDB-41143: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to one mink ACE2 receptors
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Liang B

PDB-8t20: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to two mink ACE2 receptors
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8t21: 
Cryo-EM structure of mink variant Y453F trimeric spike protein
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8t22: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to one mink ACE2 receptors at downRBD conformation
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8t23: 
Cryo-EM structure of the RBD-ACE2 interface of the SARS-CoV-2 trimeric spike protein bound to ACE2 receptor after local refinement at upRBD conformation
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8t25: 
Cryo-EM structure of the RBD-ACE2 interface of the SARS-CoV-2 trimeric spike protein bound to ACE2 receptor after local refinement at downRBD conformation.
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Zhou B, Liang B

PDB-8taz: 
Cryo-EM structure of mink variant Y453F trimeric spike protein bound to one mink ACE2 receptors
Method: single particle / : Ahn HM, Calderon B, Fan X, Gao Y, Horgan N, Liang B

EMDB-15806: 
Cryo-EM structure of the plant 80S ribosome
Method: single particle / : Smirnova J, Loerke J, Kleinau G, Schmidt A, Buerger J, Meyer EH, Mielke T, Scheerer P, Bock R, Spahn CMT, Zoschke R

PDB-8b2l: 
Cryo-EM structure of the plant 80S ribosome
Method: single particle / : Smirnova J, Loerke J, Kleinau G, Schmidt A, Buerger J, Meyer EH, Mielke T, Scheerer P, Bock R, Spahn CMT, Zoschke R

EMDB-27943: 
9H2 Fab-poliovirus 1 complex
Method: single particle / : Charnesky AJ

EMDB-27947: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ

EMDB-27948: 
9H2 Fab-poliovirus 2 complex
Method: single particle / : Charnesky AJ

EMDB-27949: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ

EMDB-27950: 
9H2 Fab-Sabin poliovirus 2 complex
Method: single particle / : Charnesky AJ

EMDB-27951: 
9H2 Fab-Sabin poliovirus 1 complex
Method: single particle / : Charnesky AJ

PDB-8e8l: 
9H2 Fab-poliovirus 1 complex
Method: single particle / : Charnesky AJ

PDB-8e8r: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ

PDB-8e8s: 
9H2 Fab-poliovirus 2 complex
Method: single particle / : Charnesky AJ

PDB-8e8x: 
9H2 Fab-Sabin poliovirus 3 complex
Method: single particle / : Charnesky AJ

PDB-8e8y: 
9H2 Fab-Sabin poliovirus 2 complex
Method: single particle / : Charnesky AJ

PDB-8e8z: 
9H2 Fab-Sabin poliovirus 1 complex
Method: single particle / : Charnesky AJ

EMDB-15674: 
Cryo-EM structure of the plant 40S subunit
Method: single particle / : Smirnova J, Loerke J, Kleinau G, Schmidt A, Buerger J, Meyer EH, Mielke T, Scheerer P, Bock R, Spahn CMT, Zoschke R

EMDB-15773: 
Cryo-EM structure of the plant 60S subunit
Method: single particle / : Smirnova J, Loerke J, Kleinau G, Schmidt A, Buerger J, Meyer EH, Mielke T, Scheerer P, Bock R, Spahn CMT, Zoschke R

PDB-8auv: 
Cryo-EM structure of the plant 40S subunit
Method: single particle / : Smirnova J, Loerke J, Kleinau G, Schmidt A, Buerger J, Meyer EH, Mielke T, Scheerer P, Bock R, Spahn CMT, Zoschke R

PDB-8azw: 
Cryo-EM structure of the plant 60S subunit
Method: single particle / : Smirnova J, Loerke J, Kleinau G, Schmidt A, Buerger J, Meyer EH, Mielke T, Scheerer P, Bock R, Spahn CMT, Zoschke R

EMDB-24075: 
Cryo-EM structure of NTD-directed neutralizing antibody LP5 Fab in complex with SARS-CoV-2 S2P spike
Method: single particle / : Reddem ER, Casner RG, Shapiro L

PDB-7mxp: 
Cryo-EM structure of NTD-directed neutralizing antibody LP5 Fab in complex with SARS-CoV-2 S2P spike
Method: single particle / : Reddem ER, Casner RG, Shapiro L

EMDB-12256: 
Structure of the autoinducer-2 exporter TqsA from E. coli
Method: single particle / : Khera R, Xie H, Michel H

EMDB-13057: 
Structure of the AI-2 exporter family protein YdiK from E. coli
Method: single particle / : Khera R, Xie H, Michel H

PDB-7nb6: 
Structure of the autoinducer-2 exporter TqsA from E. coli
Method: single particle / : Khera R, Xie H, Michel H

PDB-7ot9: 
Structure of the AI-2 exporter family protein YdiK from E. coli
Method: single particle / : Khera R, Xie H, Michel H

EMDB-13453: 
Cryo-EM structure of the agonist setmelanotide bound to the active melanocortin-4 receptor (MC4R) in complex with the heterotrimeric Gs protein at 2.6 A resolution.
Method: single particle / : Heyder NA, Schmidt A, Kleinau G, Hilal T, Scheerer P

EMDB-13454: 
Active Melanocortin-4 receptor (MC4R)- Gs protein complex bound to agonist NDP-alpha-MSH at 2.86 A resolution.
Method: single particle / : Heyder NA, Schmidt A, Kleinau G, Hilal T, Scheerer P

PDB-7piu: 
Cryo-EM structure of the agonist setmelanotide bound to the active melanocortin-4 receptor (MC4R) in complex with the heterotrimeric Gs protein at 2.6 A resolution.
Method: single particle / : Heyder NA, Schmidt A, Kleinau G, Hilal T, Scheerer P

PDB-7piv: 
Active Melanocortin-4 receptor (MC4R)- Gs protein complex bound to agonist NDP-alpha-MSH at 2.86 A resolution.
Method: single particle / : Heyder NA, Schmidt A, Kleinau G, Hilal T, Scheerer P

EMDB-12215: 
pre-50S-ObgE particle state 1
Method: single particle / : Hilal T, Nikolay R

EMDB-12216: 
pre-50S-ObgE particle state 2
Method: single particle / : Hilal T, Nikolay R, Spahn CMT

EMDB-12217: 
in vitro reconstituted 50S-ObgE-GMPPNP-RsfS particle
Method: single particle / : Hilal T, Nikolay R

EMDB-12218: 
pre-50S-ObgE particle
Method: single particle / : Hilal T, Nikolay R, Spahn CMT, Schmidt S

EMDB-12219: 
50S-ObgE-GMPPNP particle
Method: single particle / : Hilal T, Nikolay R

PDB-7bl2: 
pre-50S-ObgE particle state 1
Method: single particle / : Hilal T, Nikolay R, Schmidt S, Spahn CMT

PDB-7bl4: 
in vitro reconstituted 50S-ObgE-GMPPNP-RsfS particle
Method: single particle / : Hilal T, Nikolay R, Spahn CMT

PDB-7bl6: 
50S-ObgE-GMPPNP particle
Method: single particle / : Hilal T, Nikolay R, Schmidt S, Spahn CMT
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
